Research overview
・A rule table of human cells
Our lab was founded in 2012 for the purpose of bringing an informatics approach to the study of iPS cells. After spending two years laying done the infrastructure needed, including that to operate the iPS Cell Stock at CiRA, we began our theoretical cell analysis. Our goal is to generate a rule table of human cells on a human cell database, in which we can objectively compare cell states and elucidate cell evolution. Along with conducting the theoretical experiments needed, we also conduct wet experiments to verify our theory.

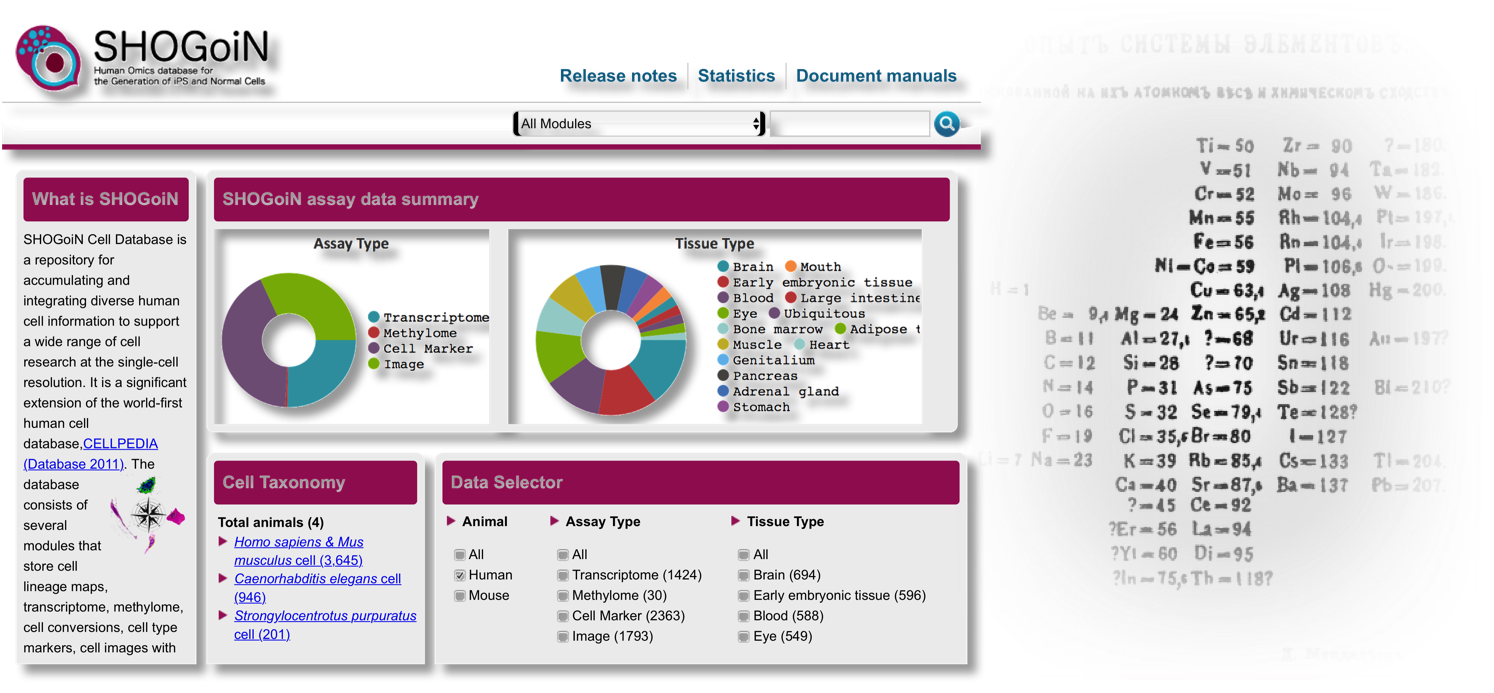
CiRA has established protocols to induce a wide range of cell types from iPS cells. To ensure the safety of these cells for clinical application, we need strict and robust checks. We are developing single-cell technologies for this purpose. (1) Currently, we have examined the gene expressions of 96 iPS and ES cells at the single-cell level simultaneously. In conjunction with this work, we reported SHOGoiN, a database to store our results. This database is intended for quality control when preparing cells for regenerative medicine.
There are currently more than 20 stem cell banks in the world, but currently no standard provisions for preparing and comparing cells between these banks exists. We are therefore working with international consortia to define Minimum Information Standard (MIACARM) for this purpose.
・In silico three-dimensional representation of human tissue
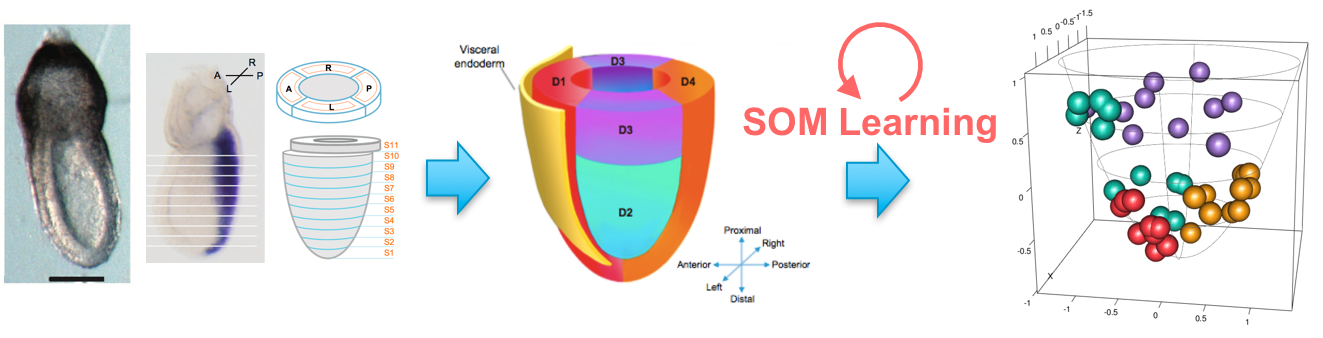
Based on the single-cell gene data in SHOGoiN, we have developed an algorithm to efficiently extract gene expression information regarding human cell differentiation. Recent gains in technology have allowed for the analysis of massive gene expression data. Using only these data and applying numerical and statistical methods, we can extract at the single-cell level which genes are essential for differentiation into particular cell types. We are integrating these gene expression data with data on methylation levels and other omics to reconstruct three-dimensional human tissue in silico. (2)
・Cell therapy and toxicity evaluation system using AI and machine learning
In 2016, we published a paper on a chemical toxicity prediction system using human ES cells (3), and established 、“Stem Cell-based Chemical Risk Information Sharing Consortium (scChemRISC)”. Through the consortium, we are developing a risk evaluation system using machine learning and AI technologies not only for drug discovery but also for safety evaluation in food and chemical industries.
(Reference: The Kyoto Shimbun (Oct. 28, 2017, morning edition P27), The Mainichi (Oct. 28, 2017, morning edition P26)
(1) Panina et al. Exp. Mol. Med. (2020)
(2) Mori et al. Scientific Rep. (2019)
(3) Yamane et al. Nucleic Acids Res. (2016)